12 Automating Pipelines: RiFA – Resistance in Plasmodium falciparum Amplicon Sequences
RiFA (Resistance in Falciparum Amplicon) is a bioinformatics workflow designed to detect drug resistance mutations in targeted amplicon sequencing data from Plasmodium falciparum, the primary cause of severe malaria. It focuses on key genes linked to resistance against major antimalarial drugs such as artemisinin, chloroquine, sulfadoxine-pyrimethamine, and others.
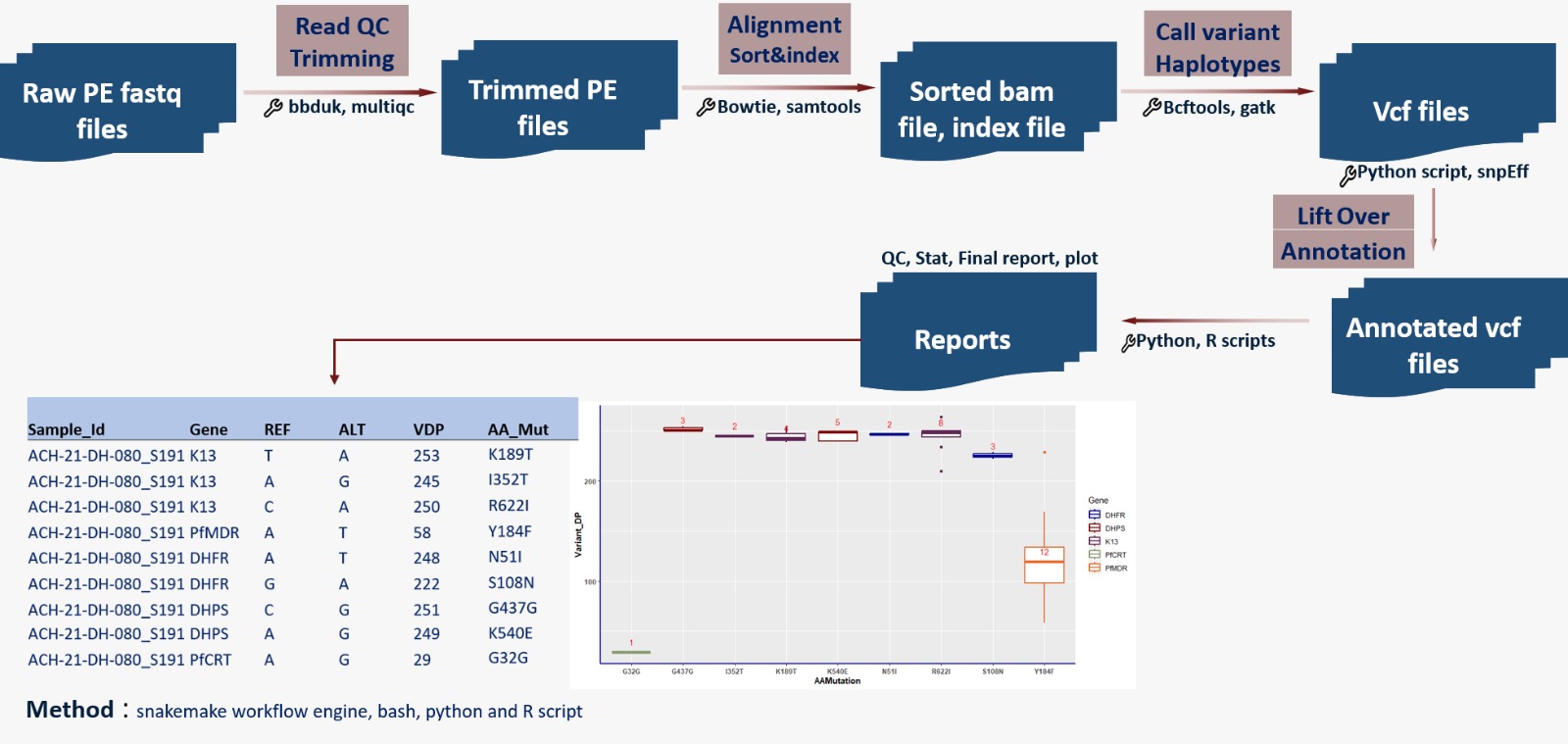
This pipeline automates processing of amplicon reads → quality control → alignment → variant calling → resistance mutation annotation — making it efficient, reproducible, and scalable for molecular surveillance of antimalarial resistance.
12.1 What Are Workflow Pipelines?
A workflow pipeline is a sequence of automated computational steps that process and analyze data in a structured, reproducible way.
12.1.1 Why Pipelines Matter in Bioinformatics
- Efficiently handle large datasets (hundreds/thousands of samples)
- Minimize manual errors and inconsistencies
- Enable reproducible research — run the same analysis tomorrow or years later with identical results
- Automate complex multi-step analyses that would otherwise be time-consuming
12.1.2 Classic Amplicon Sequencing Pipeline Example
12.2 Popular Workflow Managers / Engines
- Snakemake — Python-based, rule-based, great for dependency management
- Nextflow — DSL-based, highly portable (local, HPC, cloud)
Workflow managers help you:
- Define clear steps and their dependencies
- Execute tasks automatically in the correct order
- Parallelize jobs across cores / nodes to accelerate analysis
- Handle software environments via Conda, Docker, or Singularity
12.2.1 Where Can Pipelines Run?
- Personal laptop / workstation
- High-Performance Computing (HPC) clusters
- Cloud platforms (AWS Batch, Google Cloud Life Sciences, Azure Batch)
12.3 Key Benefits of Workflow Pipelines
Efficiency
Automates repetitive tasks → frees researchers for interpretation rather than clicking.
Reproducibility
Version-controlled pipelines ensure others (or future you) can reproduce results exactly.
Scalability
Process 10 samples or 10,000 samples with minimal changes.
Collaboration & Sharing
Share via GitHub, GitLab, or Zenodo → colleagues worldwide can reuse, adapt, and cite your work.
12.4 Hands-on Exercise: Run the RiFA Pipeline
In this exercise, you will use the RiFA pipeline to identify drug resistance mutations in Plasmodium falciparum amplicon sequences.
12.4.1 Target Genes in RiFA
RiFA focuses on key resistance-associated loci:
- pfcrt — chloroquine resistance transporter
- pfmdr1 — multidrug resistance protein 1
- pfk13 (kelch13) — artemisinin partial resistance
- pfdhfr — pyrimethamine resistance
- pfdhps — sulfadoxine resistance
- cytb — atovaquone resistance (cytochrome b)
12.4.2 Pipeline overview
12.4.3 Getting Started
Visit the official repository:
https://github.com/BettyAC/RiFA
→ Read the full README, installation guide, and example usage.Clone the repository:
git clone https://github.com/BettyAC/RiFA.git cd RiFACreate and run the environment
# Create and activate the environment (file name may vary — check README) conda env create -f environment.yml conda activate rifa-envRun the pipeline
snakemake --use-conda --cores 8 --rerun-incomplete
Expected Outputs:
- Annotated variant table (CSV/TSV) with resistance mutations, allele frequencies, and quality metrics
- MultiQC report summarizing QC and coverage
- Visual summaries (coverage plots, mutation heatmaps, frequency bar charts)
- Resistance profile summary per sample
Note: This pipeline is part of efforts like the Ethiopian Malaria Genomics Network (EMAGEN) and similar surveillance projects. Always check the repository for the latest version, example datasets, and any required reference files.